|
We present protocols on how to perform multiplex hextuple luciferase in the accompanying Current Protocols in Molecular Biology article (see Sarrion-Perdigones, et al., in press). Protocols are provided on how to prepare DNA components and destination vector plasmids, design synthetic DNA, and perform assembly cloning of new transcriptional reporter elements, implement multipartite synthetic assembly cloning of single pathway luciferase reporters, and carry out one step assembly of final multiplex hextuple luciferase vectors. This improved assembly protocol provides opportunities to analyze any five desired pathways at once much quicker. Here we present an improved three-step synthetic assembly protocol to quickly and efficiently generate multiplex hextuple luciferase reporter plasmids for other signaling pathways of interest. The same synthetic assembly cloning pipeline allows the stitching of numerous other cellular pathway luciferase reporters. Our proof-of-concept multiplex hextuple luciferase assay was designed to simultaneously monitor the p53, TGF-β, NF-κβ, c-Myc, and MAPK/JNK signaling pathways. Because all six reporters are on a single piece of DNA, a single vector ensures stoichiometric ratios of each transcriptional unit in each transfected cell, resulting in lower experimental variation. We used synthetic assembly cloning ( Vazquez-Vilar, et al., 2020) to assemble all six luciferase reporter units into a single vector, over eight stitching rounds. We recently developed a multiplex luciferase assay that allows monitoring the activity of five experimental pathways against one control simultaneously. They are genetically encoded, versatile, and cost-effective, whose output signals can be sensitively detected. Luciferases are good candidates for generation of such signals. The signals from such measurements should be independently detectable and measure large dynamic ranges. While most cell-based assays measure single quantities, multiplexed assays seek to address these limitations by obtaining multiple simultaneous measurements. Such assays are designed to examine the effects of small compounds on targets, pathways, or phenotypes participating in normal and disease processes. Planning, visualization, and documentation software for gene analysis in molecular biology methods that support the range of cloning and PCR operations.Make the DNA constructs created in your research an easy-to-read electronic document.These document files can be shared among researchers using the free viewer (SnapGene Viewer).High-throughput cell-based screening assays are valuable tools in the discovery of chemical probes and therapeutic agents. – Automatically record all operations in the cloning procedure – Visualization of DNA and chromosomes (chromosome size sequences) – Simulation of PCR and common cloning methods Gsl biotech snapgene viewer software# Addition and removal of restriction enzymes by silent mutation – Convert various file formats in molecular biology to SnapGene – Automatically create change history with excellent identification such as color coding Data management with Data Retention & Folder Self-Service explained.– Plasmid sequence and annotation, alignment, agarose gel electrophoresis Any feature type or qualifier can be defined (added / deleted) The silent mutation enzyme site tool allows you to easily add or remove restriction enzyme sites from your coding sequence (you can add / remove restriction sites to your coding sequence without changing the translation).Ģ. 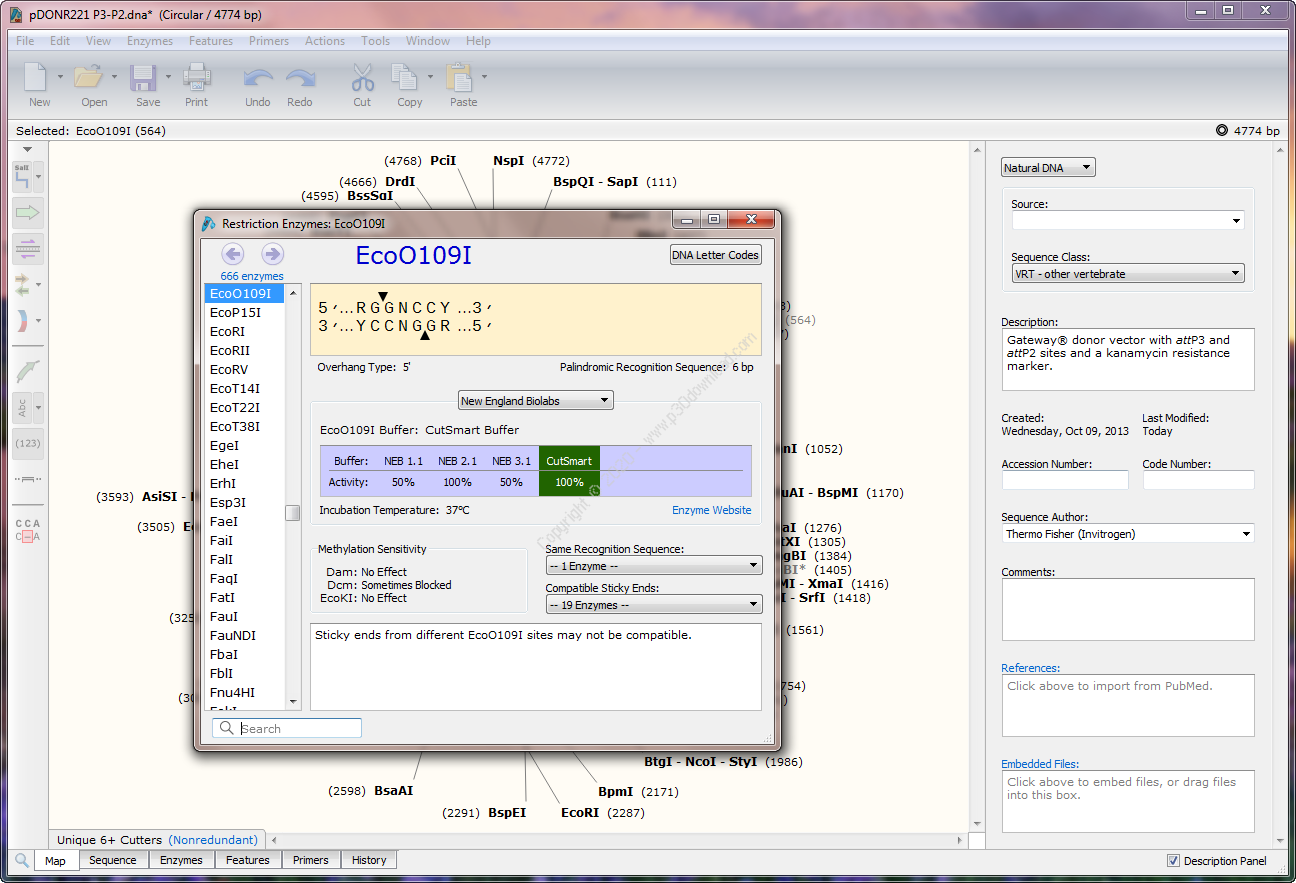
It is now possible to handle any feature type other than the preset feature type (GenBank).In addition to adding and removing feature types, you can also change the default feature color to help identify.ģ. Agarose gel simulation can be saved as a. You can now save the agarose gel simulation as a. Functional annotation in pairwise and multiple alignment gel file, share the gel file with other users, view it, or view and edit the gel after saving.Ĥ. You can view, add, and edit features in both pairwise and multiple sequence alignments.You can also switch between the Alignment view and the Features view to see the sequence, features, modifier information associated with each feature, and more.ĥ. The cloning simulation dialog has been revamped for greater flexibility.You can add, remove, and reorder fragments within the cloning interface.Also, in the simulation of restricted cloning, Gibson Assembly, In-Fusion Cloning, and NEBuilder HiFi Assembly, it is possible to set the "Orientation of Vector" of the vector in the opposite direction if necessary. ■ Click here for product details and inquiries * For other functions, Release notes for SnapGene 6.0 has more details.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
